SNVPhyl: Whole Genome SNV Phylogenomics Pipeline
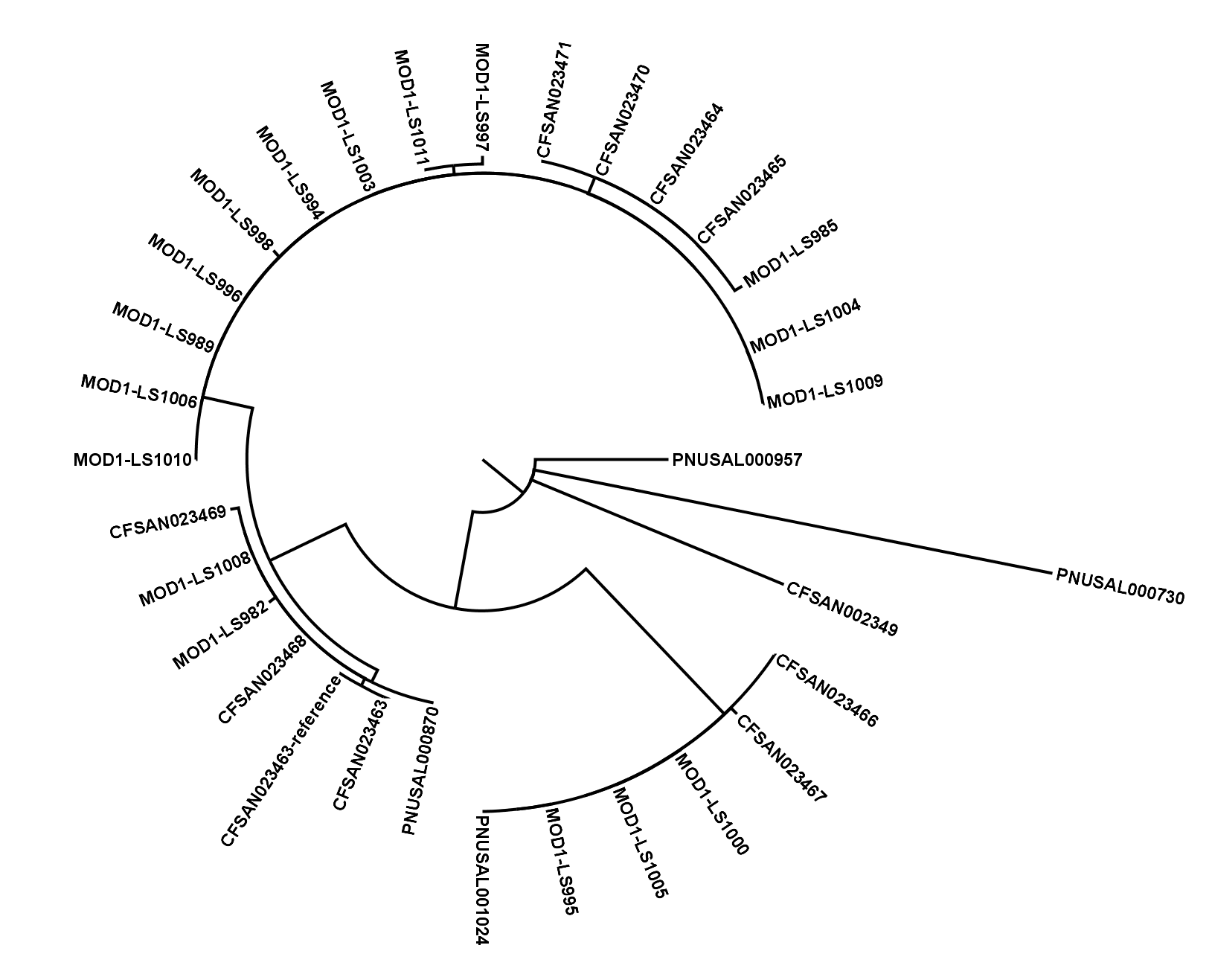
The SNVPhyl (Single Nucleotide Variant PHYLogenomics) pipeline is a pipeline for identifying Single Nucleotide Variants (SNV) within a collection of microbial genomes and constructing a phylogenetic tree. Input is provided in the form of a collection of whole genome sequence reads as well as an assembled reference genome. The output for the pipeline consists of a whole genome phylogenetic tree constructed from the detected SNVs, as well as a list of all detected SNVs and other information.
Operation
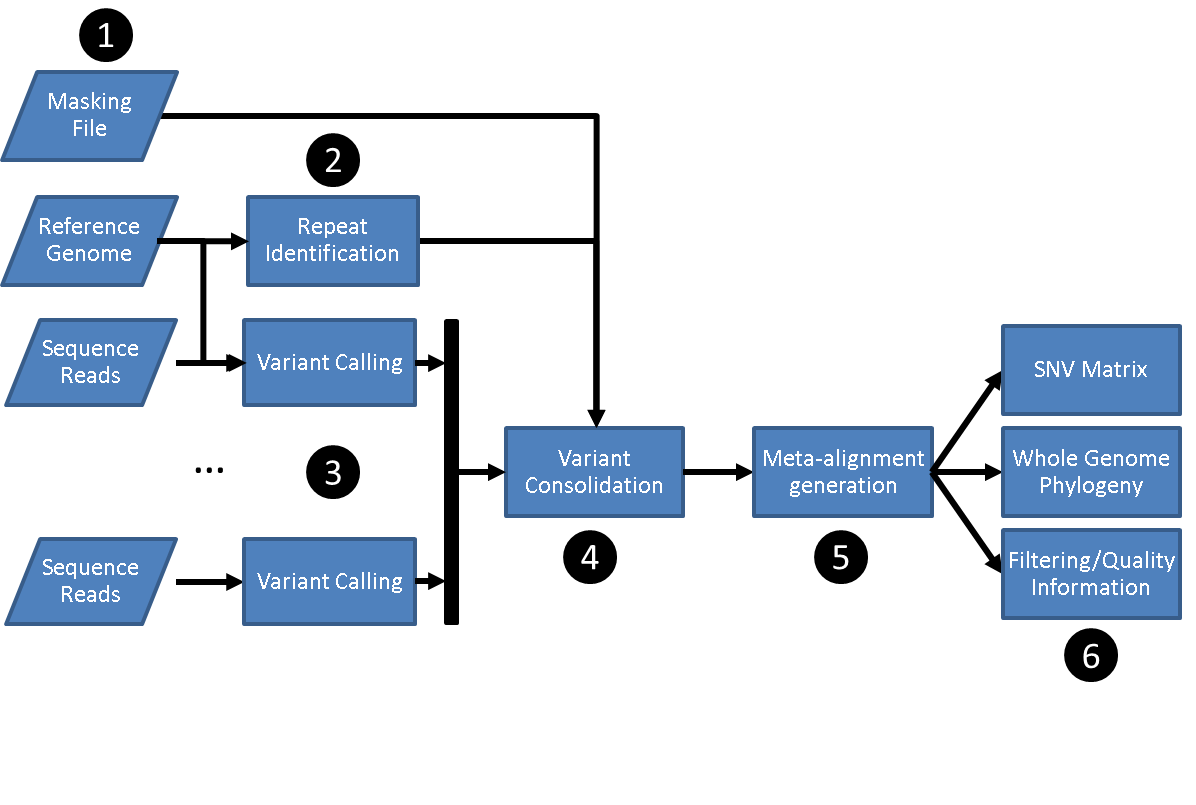
SNVPhyl identifies variants and generates a phylogenetic tree by mapping the input sequence reads to a reference genome followed by filtering out any invalid variant calls. The stages are as follows:
- Preparing input files including:
- A set of sequence reads.
- A reference genome.
- An optional file of regions to mask on the reference genome.
- Identification of repeat regions on the reference genome using MUMMer.
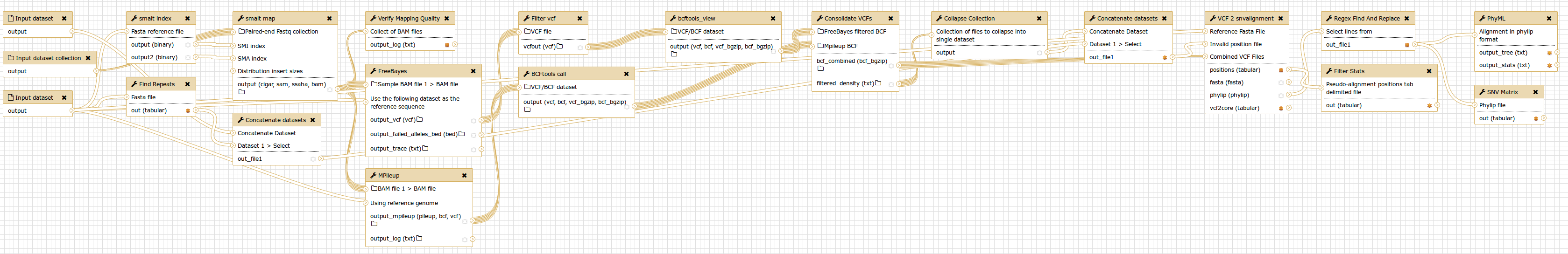
- Reference mapping and variant calling using SMALT, FreeBayes and SAMtools/BCFtools.
- Merging and filtering variant calls to produce a set of high quality SNVs.
- Generating an alignment of SNVs.
- Building a maximum likelihood tree with PhyML and generating other output files.
SNVPhyl is implemented as a Galaxy workflow, with each of these stages implemented using a specific Galaxy tool.
More information on the operation and installation of the pipeline can be found in the Usage and Installation sections.
Code is available on GitHub under the https://github.com/phac-nml/snvphyl-galaxy, https://github.com/phac-nml/snvphyl-tools, and https://github.com/phac-nml/snvphyl-galaxy-cli projects.
Contact
Comments, questions, or issues can be sent to Aaron Petkau - aaron.petkau@phac-aspc.gc.ca.